New Details about Hepatitis C Protein May Aid Development of Vaccine
Scientists have mapped the structure of a hepatitis C virus surface protein that is responsible for binding to host liver cells, leading to virus entry.
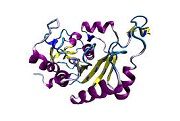
Scientists have figured out a protein structure of the hepatitis C virus that could bring researchers closer to developing a vaccine to prevent the disease.
Research published online in the “Letters” section of the journal Nature describes the viral structural protein E2, which helps the hepatitis C virus (HCV) evade the body’s natural immune system response, said Joseph Marcotrigiano, an author of the study and professor of chemistry and chemical biology at Rutgers University in New Jersey. The research resulted from a collaboration of scientists from Rutgers’ Center for Advanced Biotechnology and Medicine and the Emory University School of Medicine.
There currently is no vaccine for HCV. Finding out more about the structure of the virus could bring scientists closer to creating one, Marcotrigiano said.
“Viruses are smart and it is a constant battle to keep them out,” said Marcotrigiano in a news release that accompanied publication of the study results. “That’s why the development of a vaccine is so important. It’s always better to prevent infection through an effective vaccine then to treat after a chronic infection has been established.”
Hepatitis C causes inflammation of the liver, can lead to cirrhosis of the liver, and is the leading cause of liver transplants in the United States. The Centers for Disease Control estimates that 3.2 million people in the U.S. are infected with HCV. Though it is the most common chronic blood-borne infection, many people don’t know they have it because symptoms may not appear until the virus has severely damaged the liver.
Hepatitis C mutates constantly, which allows the virus to infect a host cell and dodge immune responses, while spreading infection throughout the body. Two envelope glycoproteins, E1 and E2, are on the outside membrane of the virus and are responsible for binding to host liver cells, leading to virus entry. Knowing more about the E2 protein structure could assist with future research aimed at understanding the binding mechanism of HCV.
“For many other viruses, like influenza, the entry pathway is well worked out. But in HCV, there is much we do not yet understand,” Marcotrigiano said.
Previously it was thought that the HCV E2 protein was a member of what is known as the class II viral fusion proteins found in related viruses such as yellow fever, Dengue and West Nile. In fact, this new research reveals that E2 has very little in common with these proteins.
“So E2 has some similarities, but is actually very different than class II fusion proteins,” Marcotrigiano said.
The Nature article describes the E2 core as a compact, globular domain structure, consisting mostly of β-strands and random coil with two small α-helices. The fold consists of many strands that are held together by an extensive hydrophobic core and disulphide bonds. Marcotrigiano explained that Sheet A has an IgG-like fold commonly found in viral and cellular proteins but sheet B represents a novel fold, possibly unique to HCV.
Knowledge of viral protein structures has been critical to development of new drugs. Finding out more about the structure of HCV could bring scientists closer to creating a vaccine to prevent the disease, Marcotrigiano said.
“Structure gives you a foundation for further studies… an unusual new look at basically how things work,” explained Marcotrigiano.
