HIV Structural Discoveries Could Lead to Advanced Treatments
“HIV is a clever virus and has learned to evade even some of the best drugs on the market," said Dmitry Lyumkis, PhD.

A team from Salk Institute in La Jolla, California solved a piece of the human immunodeficiency virus (HIV) puzzle that’s been stumping researchers for decades.
HIV establishes itself in human DNA—which is a sign that the disease is progressing—and replicates in the body. While there is no cure, antiretroviral therapy can significantly slow the progression and improve patients’ lives. The Salk Institute scientists uncovered the atomic structure on a piece of machinery which assists in that HIV-DNA integration.
“HIV is a clever virus and has learned to evade even some of the best drugs on the market. Understanding the mechanisms of viral escape and developing more broadly applicable drugs will be a major direction in the future,” senior author, Dmitry Lyumkis, PhD, said in a news release.
HIV takes a DNA copy of its RNA genome and inserts it into human DNA by using the intasome—the piece of machinery mentioned earlier. The intasome uses enzymes, called integrases, to contribute to the viral DNA. Researchers discovered structural information on a part of the integrase enzyme back in 1994, but conventional techniques have left questions about the whole structure of the intasome.
One HIV treatment option is a class of drugs called integrase strand transfer inhibitors (INSTIs). The drugs target the intasome, but lingering mechanism questions have left researchers wondering how they exactly work against the virus.
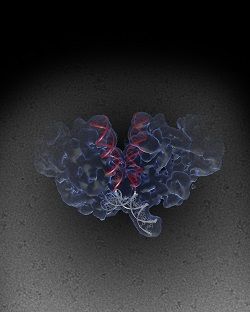
Using a new imaging technique called single-particle cryo-electron microscopy (cryo-EM), the Salk Institute team was able to increase the molecule images. “The team attached a specific protein to improve the intasome’s ability to dissolve in liquid and bathed the intasome in a syrup-like liquid called glycerol, with loads of salt added to prevent it from clumping,” the statement explained. It was noted that this was an extreme circumstance for the cryo-EM, but necessary for the HIV intasome.
“We’re particularly excited about the ability to understand and combat mechanisms of viral resistance,” Lyumkis continued. “Now we have the very first native blueprint in the context of HIV for studying the mechanisms of INSTIs.”
All retroviruses, such as prototype foamy virus (PFV) and human T cell leukemia virus, insert their RNA strand into human DNA. However, the findings indicated differences in enzyme cores of HIV and PFV. “Although these variations are minor, they could be a big deal for drug development and for understanding mechanisms of drug resistance,” Dario Passos, PhD, a senior research associate in Lyumkis’ lab. The researchers went on to say that HIV intasomes appeared more complex than other retroviruses.
Looking ahead, the hope is that this structural research will help with the development of new HIV drugs.
The study, “Cryo-EM structures and atomic model of the HIV-1 strand transfer complex intasome,” was published in the journal Science. The news release and images were provided by Salk Institute.
Related Coverage:
First Large Trial for Long-Lasting Injectable PrEP Gets Underway
Growing HIV Drug Resistance Justifies Treatment Change
New Technology Identifies Key Genes to Avoid HIV Treatment Resistance