Disease Model Links Molecular Drivers of Melanoma With Diagnostic Tests and Therapies
At the same time that the incidence of melanoma is rising globally,1 this most aggressive skin cancer is being unwound at a molecular level.
A
t the same time that the incidence of melanoma is rising globally,1 this most aggressive skin cancer is being unwound at a molecular level. Evidence has converged around the concept of driver mutations acting within molecular pathways known to play key roles in the onset, progression, and prognosis of the disease. Melanoma is coming to be understood as a highly variable form of cancer, with many subtypes that differ from patient to patient, and therapeutic outcomes dependent on matching precise subtypes to medical interventions.
Reasoning that knowledge of the genetic basis of melanoma has advanced enough to permit a classification of tumors into molecular subtypes, and that a burgeoning biotech industry is bringing the world of personalized medicine into view, a group of industry and academic experts collaborated to create a Melanoma Molecular Disease Model.2 Published in 2011, the model links melanoma molecular pathways with key genes and biomarkers, available diagnostic technologies, and potentially effective therapies.
The concept behind the Melanoma Molecular Disease Model was demonstrated in the summer of 2011, with the approval of vemurafenib (Zelboraf, Genentech) to treat patients with metastatic melanoma tumors that express the BRAFV600E mutation. The drug was licensed on the basis of phase III trial evidence showing an improvement in overall survival,3 and was approved together with the Cobas 4800 BRAF V600 Mutation Test to identify the BRAF mutation.4 (See “New Therapies Emerge for Metastatic Melanoma.”)
Developing the Model
Early in the process of developing the Melanoma Molecular Disease Model, Vidwans and colleagues assembled a panel of recognized melanoma experts whose consensus judgment resulted in the initial map of melanoma subtypes. Each subtype comprising the initial list is defined by an association with a key oncogene/tumor suppressor (eg, BRAF, cKIT), that either acts alone or in combination with other oncogenes that play supportive roles.2
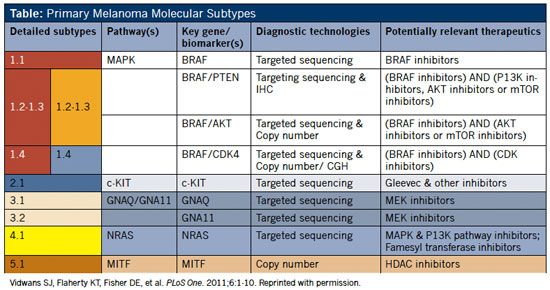
The model’s principal molecular subtypes were then ranked in rough order of importance based on prevalence among patients, as well as the current potential for targeted therapeutic intervention. Variations under major subtypes (eg, 1.1, 1.2) reflect a recognition that a melanoma may contain aberrations in more than one pathway (Table, page 2).2
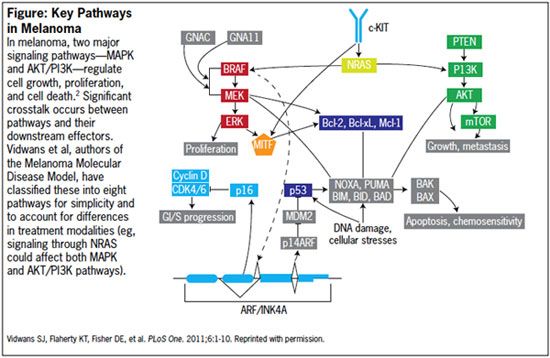
Figure: Key Pathways in Melanoma
Pathways
Rollover to enlarge.
The MAPK Pathway
The mitogen-activated protein kinase (MAPK) pathway is especially important to melanoma tumorigenesis and regulation of cell growth, proliferation, and differentiation. Genetic aberrations in the MAPK pathway are reportedly found in over 80% of cutaneous melanomas, with abnormalities in RAS, RAF, MEK and ERK.2,5 Melanoma molecular subtype 1 is defined overall by aberrations in the MAPK pathway, either alone or in combination with other pathways (eg, AKT/PI3K, CDK).2
Melanoma Subtype 1.1
The most common mutation in the MAPK pathway (melanoma subtype 1.1) is in the BRAF gene.2 This mutation occurs in 40% to 70% of melanoma cases.6 In addition to the recently approved vemurafenib, another BRAF inhibitor, sorafenib, has been investigated in phase I, II, and III clinical trials.2,7 Results to date suggest that sorafenib as monotherapy or in combination with chemotherapy may be of limited use,7 or the subpopulation of patients with advanced melanoma that will receive benefit from sorafenib awaits identification of the right biomarker.2,7
Melanoma Subtypes 1.2 & 1.3
Mounting evidence suggests that melanoma development and metastasis may require multiple pathways. Accordingly, melanoma subtypes 1.2 and 1.3 define cases that possess abnormalities in both the MAPK and AKT/PI3K pathways. Some drug development initiatives for these subtypes call for combination therapy, with inhibitors of both pathways.2
Melanoma Subtype 1.4
Subtype 1.4 is associated with mutations in both the MAPK and CDK pathways, resulting in activation of BRAF and overexpression of CCND1/Cyclin D. (The CDK pathway has been proposed as a contributor to metastasis in melanoma with BRAF mutations.)2
The rational approach to therapy for the melanoma subtype 1.4, according to the authors of the Melanoma Molecular Disease Model, is a combination of CDK and BRAF inhibitors.2
Melanoma Subtype 2 Overview
Subtype 2 is characterized by mutations in the c-KIT pathway, a receptor tyrosine kinase (RTK) involved in intracellular processes such as cell growth, cell division, and migration in response to Stem Cell Factor (SCF) activity.2 Several approved drugs target c-KIT, including imatinib, sunitinib, nilotinib, and dasatanib.2
Melanoma Subtype 2.1
As of the publication of the Melanoma Molecular Disease Model in 2011, c-KIT is the only subtype in this category. It is characterized by genetic aberrations in c-KIT, which generally include mutations and/or copy number increases. One study reported the occurrence of these mutations in 39% of mucosal, 36% of acral, and 28% of melanomas on chronically sun-damaged skin (but not in melanomas on skin without chronic sun damage).2
c-KIT inhibitors represent the rational treatment strategy for this subtype of melanoma, and clinical evidence has provided support for the efficacy of therapies such as imatinib, sorafenib, temozolamide, and dasatinib.2
One example of the potential of a precise matchup of mutation to target drug was demonstrated in a study by a group at MD Anderson Cancer Center. The group had previously succeeded in identifying the first human melanoma cell line with L576P, the most common KIT mutation in melanoma.8
After determining that the mutation was not responsive to KIT inhibitors imatinib, nilotinib, or sorafenib, but was responsive to dasatinib in the laboratory, the researchers treated two patients with metastatic melanoma with dasatinib. Both patients tested positive for the L576P KIT mutation, and one of the patients had previously received treatment with imatinib. Following dasatinib treatment, both patients experienced a more than 50% reduction of their tumors.8
The c-KIT inhibitor imatinib also was associated with significant activity in patients with metastatic melanoma and c-KIT mutations. In a phase II open-label, single-arm trial of 43 patients, the overall response rate was 23.3%.9
Melanoma Subtype 3, 3.1
Subtype 3 is characterized by mutations in the G proteins, GNAQ and GNA11. In subtype 3.1, a mutation in the GNAQ gene affects codon 9, which may drive constitutive activity in the MAPK pathway.2
Melanoma Subtype 3.2 Overview
Subtype 3.2 is characterized by a mutation in the GNA11 gene that affects codon 209. This mutation may drive constitutive activity of the MAPK pathway. 2 Several MEK inhibitors in development could be useful to treatment of this subtype.
Table: Primary Melanoma Molecular Subtypes
Subtypes
Rollover to enlarge.
Melanoma Subtype 4
Subtype 4 is characterized by abnormalities in the RAS gene. RAS-regulated signal pathways are involved in control of proliferation, differentiation, cell adhesion, apoptosis, and cell migration via MAPK and AKT/PI3K pathways.2
Melanoma Subtype 4.1
Subtype 4.1 is characterized by mutations in NRAS, found in about 20% of melanomas.2 (Clinical testing of an approach of concurrently targeting MAPK and AKT/PI3K pathways is planned by two pharmaceutical companies.)
Melanoma Subtype 5, 5.1
Subtype 5 is characterized by abnormalities in the melanoma development and survival pathway. In this pathway, melanocyte transcription factor (MITF—microphthalmia-associated transcription factor) is a regulator of the development, differentiation, and maintenance of melanocytes.2
Subtype 5.1 features aberrations in MITF; MITF may be an amplified locus in melanoma, and MITF amplification has been correlated with decreased survival and increased resistance to chemotherapy.2
MITF expression has been reduced by histone deacetylase (HDAC) inhibitor drugs. A study to evaluate the efficacy of panobinostat (LBH589) in patients with metastatic melanoma is under way.
The Melanoma Molecular Disease Model is a living document. Its authors expect that with time and expanding basic science knowledge, the model will acquire increasing specificity and more links to targeted therapies and diagnostics. As new information is gathered, the model is to be updated online at http://mmdm.cancercommons.org/smw/index.php/A_Melanoma_Molecular_Disease_ Model.
References:
1. MacKie RM, Hauschild A, Eggermont AM. Epidemiology of invasive cutaneous melanoma. Ann Oncol. 2009;20(suppl 6):vi1-7.
2. Vidwans SJ, Flaherty KT, Fisher DE, et al. A melanoma molecular disease model. PLoS One. 2011;6:1-10.
3. Chapman PB, Hauschild A, Robert C, et al. Improved survival with vemurafenib in melanoma with BRAF V600E mutation. N Engl J Med. 2011;364:2507-2516.
4. Zelboraf [package insert]. San Francisco, CA: Genentech USA, Inc; 2011.
5. Lemech C, Arkenau H-T. Novel treatments for metastatic cutaneous melanoma and the management of emergent toxicities. Clin Med Insight Oncol. 2012;6:53-66.
6. Davies H, Bignell GR, Cox C, et al. Mutations of the BRAF gene in human cancer. Nature. 2002;417:949-954.
7. Mangana J, Levesque MP, Karpova MB, Dummer R. Sorafenib in melanoma. Expert Opin Invest Drugs. 2012;21:557-568.
8. Woodman SE, Trent JC, Stemke-Hale K, et al. Activity of dasatinib against L576P KIT mutant melanoma: molecular, cellular, and clinical correlates. Mol Cancer Ther. 2009;8:2079- 2085.
9. Guo J, Si L, Kong Y, et al. Phase II open-label, single-arm trial of imatinib mesylate in patients with metastatic melanoma harboring c-KIT mutation or amplification. J Clin Oncol. 2011;29:2904-2908.
